About
My primary research interests are in probability and statistics, and their application to science and engineering. More details can be found in my CV (February, 2018).
February 2018 - Leukaemia and Healthy Blood Production - Part 1.
Acute myeloid leukaemia (AML) is an aggressive cancer originating from blood cells. There is an unmet clinical need for more effective therapies as the cure rate of AML is only 5-15% in patients older than 60, and the mainstay of treatment has not changed significantly in the last 30 years. About 1 in 200 men, and 1 in 250 women in the UK will get AML at some stage in their life.Perhaps surprisingly, patients with leukaemia typically present to the clinic with symptoms of insufficient healthy blood cell production such as anaemia, excessive bleeding, or recurrent infections instead of overproduction of the cancerous cells themselves. As determining how the cancer arrests healthy blood production could have significant ramifications for the design and development of treatments for leukaemia, a multi-disciplinary international team undertook two related, experimentally sophisticated, studies to address this question by studying an aggressive AML model in mice that mimics the disease found in humans.
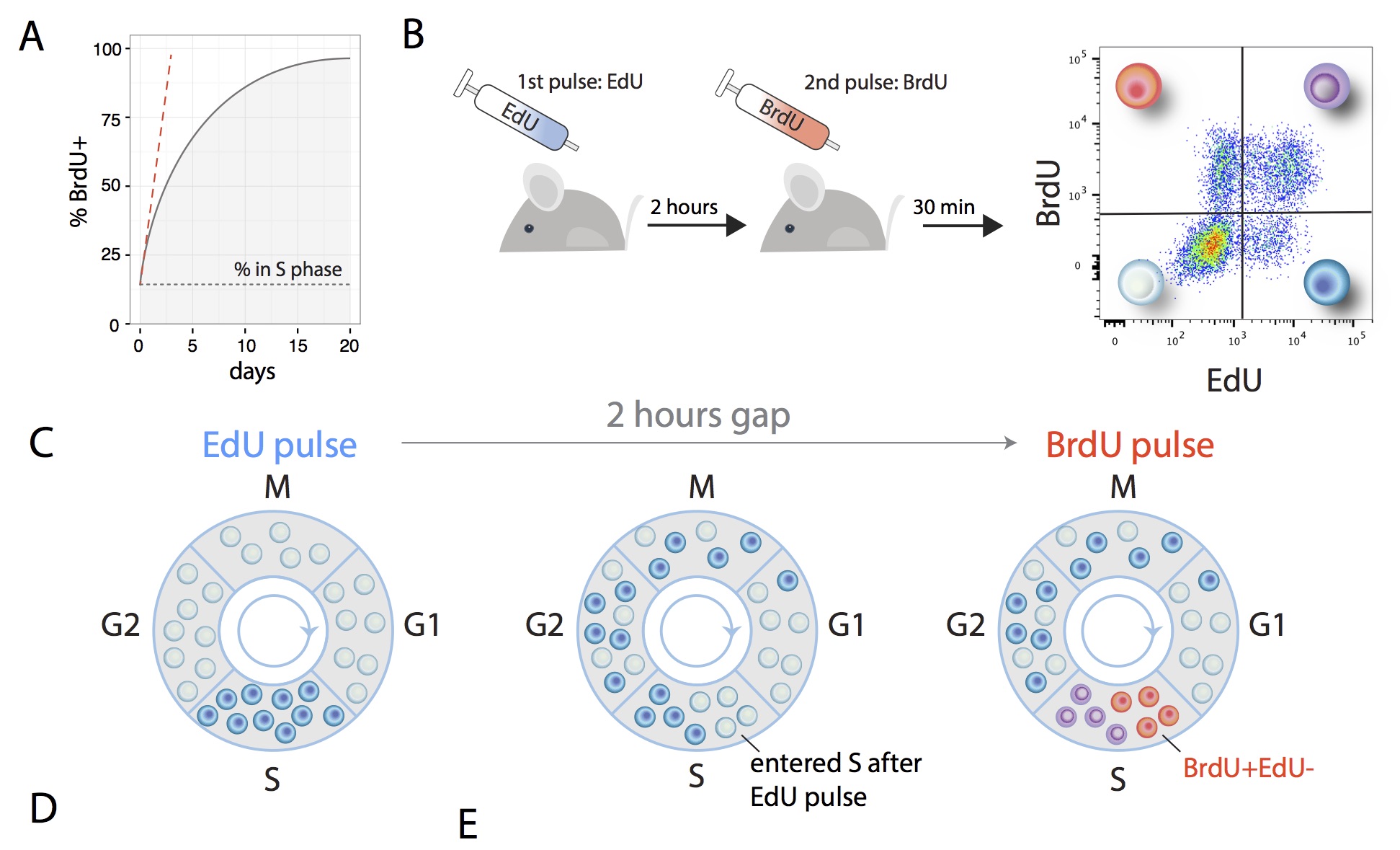
The first study was undertaken by members of Cristina Lo Celso's wet lab at Imperial College London (Olufolake Akinduro, Heather Ang, Myriam Haltalli, Nicola Ruvio, Delfim Duarte, N. M. Rashidi and Edwin D. Hawkins), in collaboration with myself and my ex-postdoc Tom Weber who is now at WEHI. To accurately determine the proliferation dynamics of both healthy blood cell production and cancer growth, as well as careful measurements of cell numbers, we implemented a method to measure the rate of cell division in vivo. To measure a rate, any applied mathematician will tell you that you need two measurements. Here the idea is that you label all cells that are currently synthesizing DNA with one label (EdU), wait a brief period and label all cells then synthesizing their DNA with a second label (BrdU) and quickly extract cells. All cells that are label positive for the second label entered DNA synthesis in the brief window between the labels, which serves as a proxy for the number of cells undergoing division per unit time.
Existing hypotheses posited a process whereby the cancer actively shuts down healthy blood production in the bone marrow, but we established that a simpler high-level mechanism was at play (Nature Communications, 2018). We found that throughout disease progression the cancer grew steadily, blithely producing new cells at a constant rate even when it ran out of space in bone marrow to host them. While the healthy cells that remained continued somewhat obliviously to the infiltration, ultimately both healthy and some cancerous cells were ousted from the bone marrow by the relentless leukaemic growth. The stem cells that are the top of the hierarchy of healthy blood cell production proved most resistant to this process, but were ultimately eliminated. En route, we identified an improved phenotypic characterization of true blood stem cells.
January 2018 - Leukaemia and Healthy Blood Production - Part 2.
While the first study, described above, provides a cellular population-level understanding of how leukaemia outcompetes healthy blood cell production, to get a grasp on why stem cells manage to resist ousting from bone marrow required distinct techniques. Cristina Lo Celso's wet lab at Imperial College London are recognised experts in intravital microscopy. That is, in the collection of optical techniques that enable visualization of biological processes in live animals. In a second study, led by her team, but encompassing a large network of researchers and clinicians spanning Glasgow to Melbourne, amongst other tools, this approach was leveraged to get to the bottom of that question.
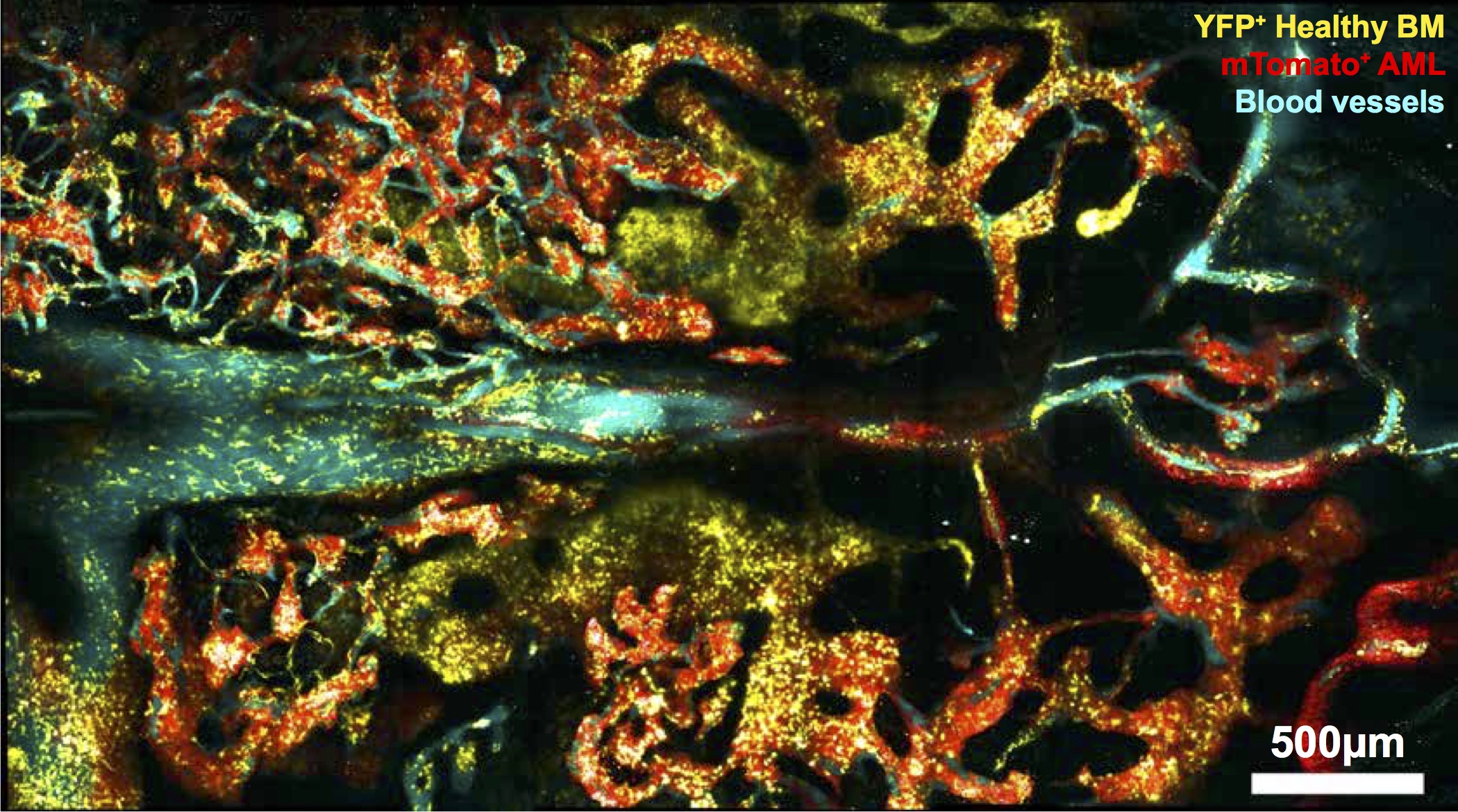
This study (Cell Stem Cell, 2018), found that leukaemia expanded in a few areas initially, gradually eliminating healthy blood cells, cells that support blood production, and the rare blood vessels normally associated with blood stem cells. By preserving those rare blood vessels with a pharmaceutical intervention it was established that healthy blood stem cells could be rescued and chemotherapeutic efficacy promoted. As the drug in question, Deferoxamine (which is commonly found in Irish clinics as a treatment for hemochromatosis), is approved for human use for a different condition, it is known to be safe. Thus, as it is already in use, it should, in principle, be possible to progress to clinical trials much quicker (~5 years) than one could with a brand new drug (~10-15 years).
Fig 1c from the paper. Mouse bone marrow showing cancerous cells in red interacting with healthy bloods cells in yellow, and blood vessels in cyan. Image created by intravital microscopy.
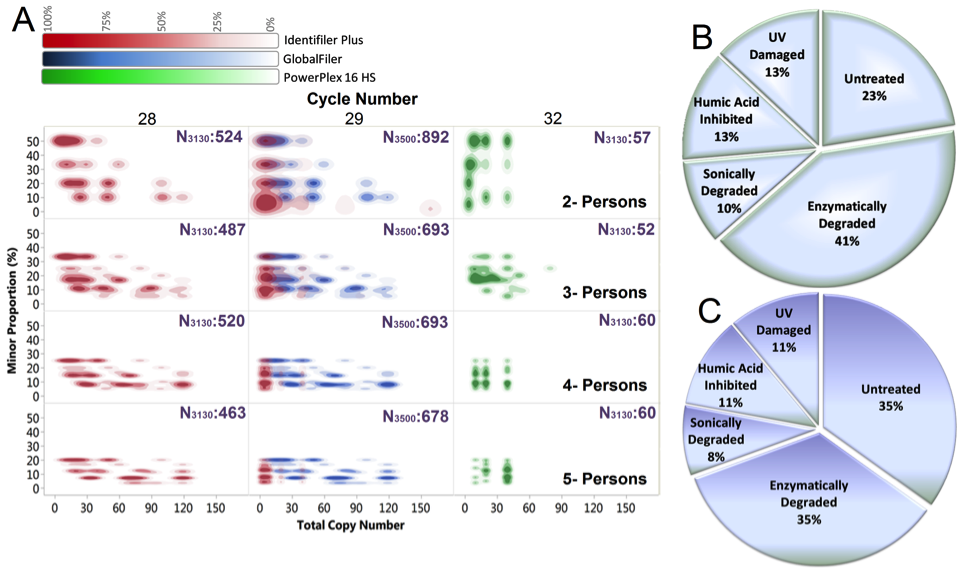
The database, which was mentioned in 2016 in a report by then US
President Barak Obama's President's Council of Advisors on Science
and Technology (PCAST), contains over 25,000 complex mixture DNA
samples created with known contributors, amounting to over six years
of experimental
work by the forensic scientist Catherine Grgicak, Lauren Alfonse and Amanda
Garrett. For these data, ground truth is known as bloods from given
genotypes were mixed and degraded to mimic crime-scene samples. These
data can be used to develop, test and confirm the accuracy new
methods for person identification by DNA. The paper describing
the database is published in
Forensic Science International:
Genetics by Lauren E. Alfonse, Amanda D. Garrett,
Desmond Lun, myself,
and
Catherine Grgicak.
October 2017 - Constructing Network Codes.
For multicast data, sending a single stream of information across a network to a range of people, Network Coding is a panacea. Creating Network Codes for non-multicast transmissions where different users want different streams, however, is remains challenging. In work led by Ying Cu, (Shanghai Jiao Tong University), with Muriel Médard, Edmund Yeh, Douglas J. Leith, and Fan Li we had a paper published in IEEE/ACM Transactions on Networking, where Communication Free Learning, an algorithm invented by Douglas J. Leith and Peter Clifford is used to construct these codes. The work leverages an old idea that Douglas J. Leith, Charles Bordenave and I had to use Doug and Peter's algorithm to solve constraint satisfaction problems, published in IEEE/ACM Transactions on Networking, 21 (4), 1298-1308, 2013.October 2017 - Network Maximal Correlation.
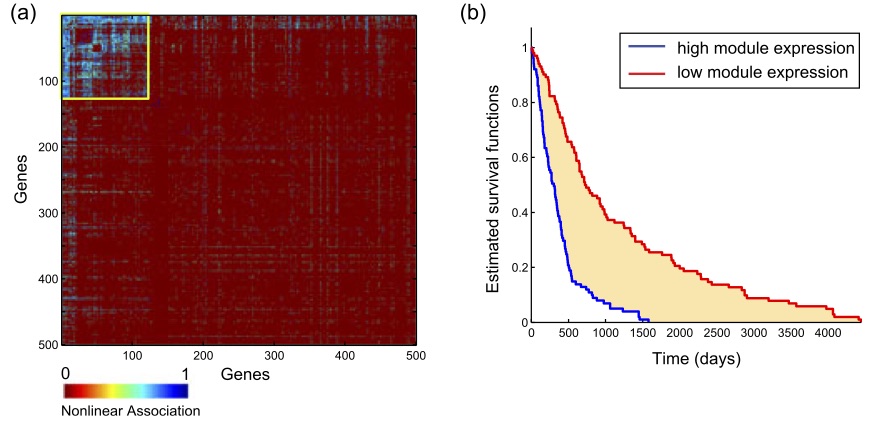
Given a pair of variables, X and Y, how do you summarize how dependent they are on each other? This is a question that is well addressed in statistics, starting with Karl Pearson's correlation coefficient, which was built on ideas that date back to work of the eugenicist Francis Galton in the 1880s. If you scatter plot X and Y, Pearson's correlation coefficient essentially summarizes the best linear description of one as a function of the other. While a useful statistic, the units in which X and Y are measured can impact the outcome. This statistic was beautifully generalised by the psychologist Charles Spearman in 1904 creating a summary of the relationship of the rank order of the data pairs, overcoming issues of units. Maximal correlation, which was introduced in the 1930s, but didn't catch on is a further generalisation that numerically summarizes a broad class of potential relationships between the variables, and there have been many more alternatives offered since then.The question of how to sumarize the correlation of more than a pair of random variables has lead to a lot of proposals and is particularly pertinent presently due to technology enabling the measurement of vast ranges of multivariate data, from the collection of genes implicated in disease to the state of traffic in a city. In a second piece of work that was undertaken with Muriel Médard and led by her students Soheil Feizi (now at Stanford), who was co-supervised by Manolis Kellis, and Ali Makhdoumi, we have had a paper published in IEEE Transactions on Network Science and Engineering with one proposal: Network Maximal Correlation. This is a natural generalisation of the largely overlooked bivariate measure, which is now tractable due to current computing power.
As an illustration of its use, it is used in the paper to identify a nonlinear
gene module of Glioma cancer that one would not find with more traditional
measures of multivariate relatedness.
March 2017 - Principal inertia components and applications.
While on sabbatical at MIT in 2015-2016, I had correlation my mind thanks to Muriel Médard's students Flávio du Pin Calmon (now at Harvard), Soheil Feizi (now at Stanford), and Ali Makhdoumi.
| All of statistics is about summary: given a set of data, how can one succinctly describe its important properties without having to relate the data set in full? One core measurement of interest is how correlated two variables are. If one measures pairs (x,y), such as (age, income), how can one efficiently report if there is a positive or negative relationship between them. One would intuitively say that the (x,y) pairs in the image to the right are positively correlated - as one gets larger, the other typically does - but you have a range of choices on how to quantify that relationship. How correlated two variables are has important ramifications for the ability to draw inferences. If two variables are correlated, perhaps you can estimate one having observed the other? This has ramifications for cases where you wish to be able to estimate one from the other such as when the pair is (what is said into the phone, what is heard over the phone), but also when you don't, such as (political preference, film you watched). |
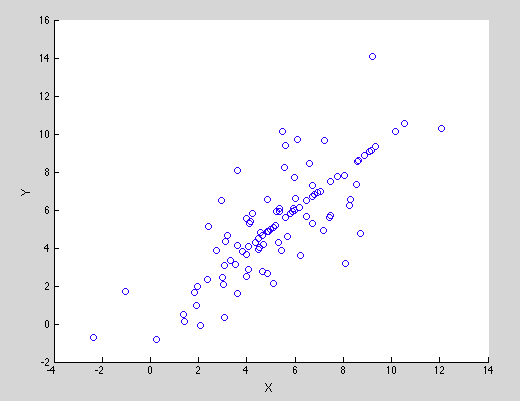
|
In work led by Flavio, published in IEEE Transactions on Information Theory, he, Ali, Muriel, Mayank Varia, my student Mark Christiansen and I explore measurements of correlation between pairs of variables taking discrete values and relate them to mechanical notions, Principal Inertia Components (PICs). The PICs lie in the intersection of information and estimation theory, and provide a fine-grained decomposition of the dependence between two variables X and Y. The PICs describe which functions of X can or cannot be reliably inferred given an observation of Y. In the paper we demonstrate that the PICs play an important role in Information Theory, while in privacy settings, the paper proves that the PICs are related to fundamental limits of perfect privacy.
January 2017 - SEEIt, A Stochastic Model of the DNA Forensics Pipeline.
With my collaborator, the forensic scientist Catherine Grgicak (BU), Kelsey C. Peters and Genevieve Wellner in her lab, and my ex-graduate student at MIT, Neil Gurram, we recently published a stochastic model that produces simulated electropherograms, the core measurement output in DNA forensics in the journal Electrophoresis. The model encodes all of the standard lab procedure, capturing sampling issues, allele drop-out, PCR amplification and stutter, and inherent instrument noise in the process. Its construction was informed and parameterized by a huge number of samples created in Catherine's lab. As those samples are costly and time-consuming to create, through its use, its main boon is in being able to quickly and cheaply create simulated mixtures with known contributors with which to explore modification of lab procedures to improve inference.
November 2016 - Clonal Cellular Programmes in the Immune System.
With my long-run immunology collaborator, Phil Hodgkin and members of his lab, Julia Marchingo, Andrey Kan, Susanne Heinzel at the Walter and Eliza Hall Institute of Medical Research, and my student Giulio Prevedello we have published a paper in Nature Communication. The article introduces a new technique that allowed us to determine the division state of offspring from individual cells, which has broad applicability, in a relatively high-throughput way that is significantly less labor intensive than microscopy methods.
The experiment design is based on a multiplex of division diluting dyes. In the early 1990s, a green fluorescent dye, CFSE, was identified by two Australian scientists, Lyons and Parish, that cells can imbibe without impacting their function. When a stained cell divides, its daughters fluoresce with half the intensity of their parent, and their daughters with half again, allowing division numbers to be determined. In the past few years, a new generation of dyes in a range of colours have been created. To enable determination of the division history of individual families, we designed a system where cells could be stained with a unique combination of colours. This enables observation of the lineages of hundreds of founding cells through multiple generations, as illustrated below with two dyes, CFSE and CTV.
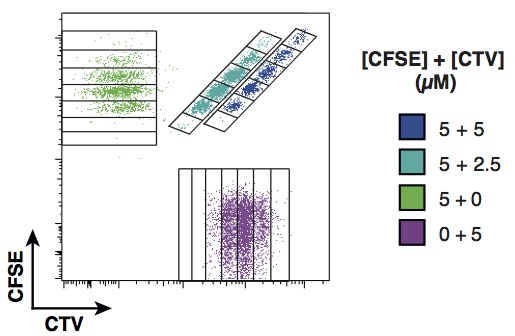
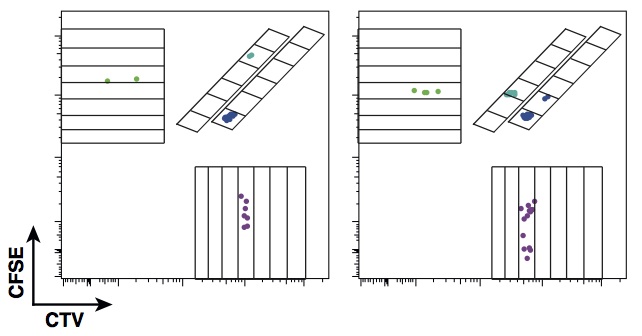
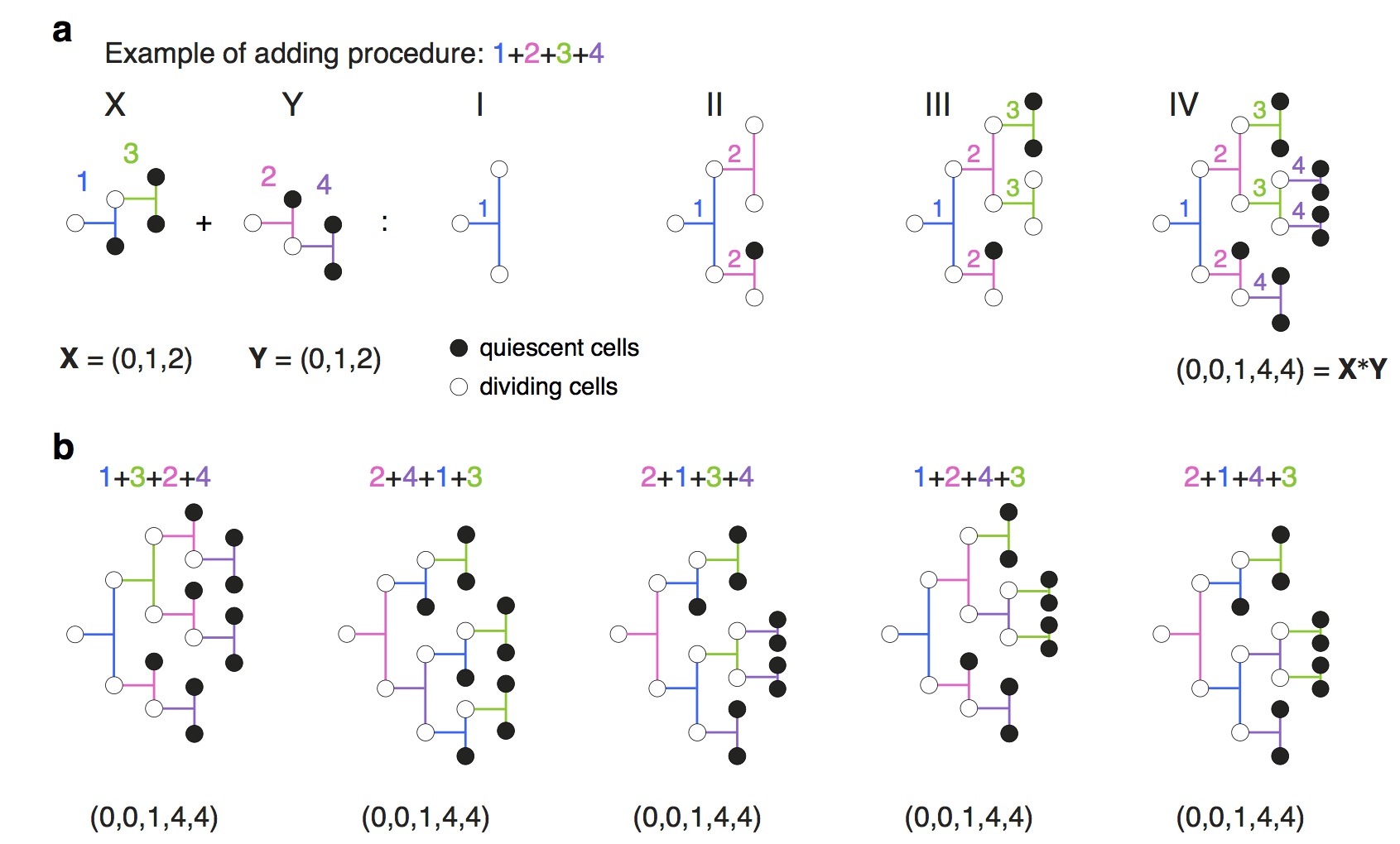
In the paper, we use the approach to consider the initial clonal expansion of T cells in response to challenge and to address the question of how individual clones integrate distinct stimuli. Contrary to expectation, the finding is that that the initial cell is being programmed independently by the stimuli to execute a regimented family expansion programme. The significant population-level heterogeneity that had been previously described comes about as a consequence of heterogeneity in the depths of individual regular trees rather than frayed family trees.
In order to address these questions and interrogate the data from a newly developed method, we had to develop some pretty new mathematics and statistics related to the concatenation of directed trees.
June 2016 - The Design of a DNA Coded Randomized Algorithm For In Situ Lineage Tracing.
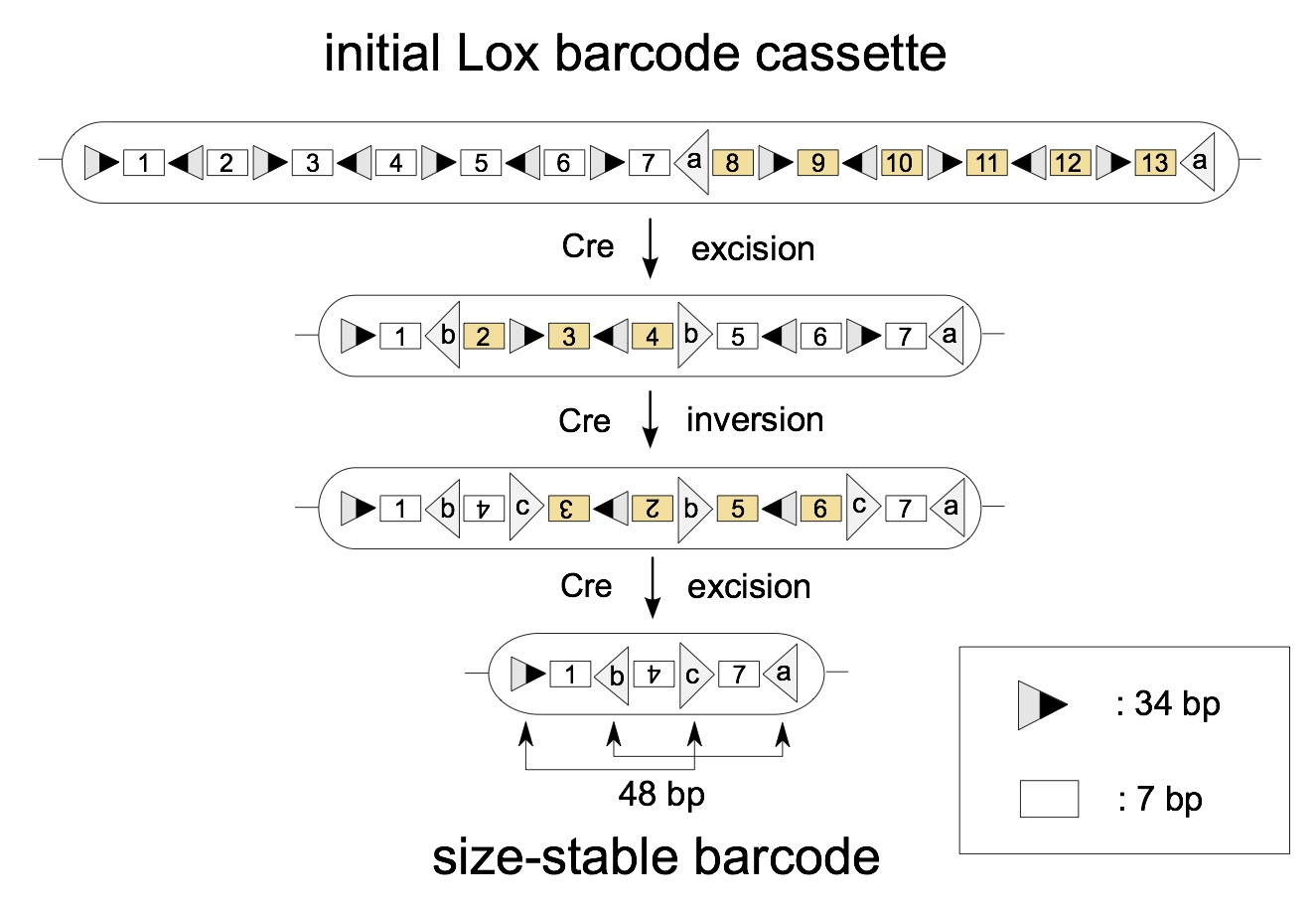
Tom Weber, our combinatorist colleague Mark Dukes from University College Dublin, and three wet lab collaborators from the Walter and Eliza Hall Institute of Medical Research, Denise Miles, Stefan Glaser and Shalin Naik, and I recently published a paper in BMC Systems Biology describing the optimal design of a randomized algorithm that can be encoded in the DNA of a cell. At an abstract level, the programme can be thought of as installing in the DNA a deck of cards, as well as mechanisms for shuffling and picking. Upon activation by the delivery of a drug, each cell executes the randomized algorithm, shuffling and removing cards from the deck until they are left with either one or three cards, whereupon the algorithm terminates.
Conceptually, the end effect is that each cell shuffles and picks either one or three cards from the deck, which are then left in that cell's DNA. As the offspring of that cell then get copies of those cards, they serve as labels them as having a shared ancestor, at least for rare combinations. Thus the algorithm's design is to maximize the diversity of the card-picks and to ensure that the selection is stable.
Starting with a deck of 13 cards, one execution of the algorithm would look as follows:
January 2016 - The Design of a DNA Coded Randomized Algorithm For Inferring Tree Depth.
How many cell division take place between birth and the construction of a brain? Are there more cell divisions between the development of skin than gut? Perhaps surprisingly, these are questions that one cannot answer at present. In a paper published in the Journal of Mathematical Biology by Tom Weber, Leïla Perié and myself, we have established a formula, which I personally think is remarkable, that may allow us to answer these questions:
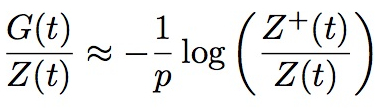
The core of the mathematics is based on newly established properties of the fundamental model of growing trees, the Bellman-Harris process first studied in 1948, which assumes substantial statistical homogeneity. We do, however, also provide a derivation of a weaker form of the formula in broad generality and establish through simulations its accuracy and utility for trees that are far from samples of a Bellman-Harris process, such as the early development of C. elegans , as illustrated here:
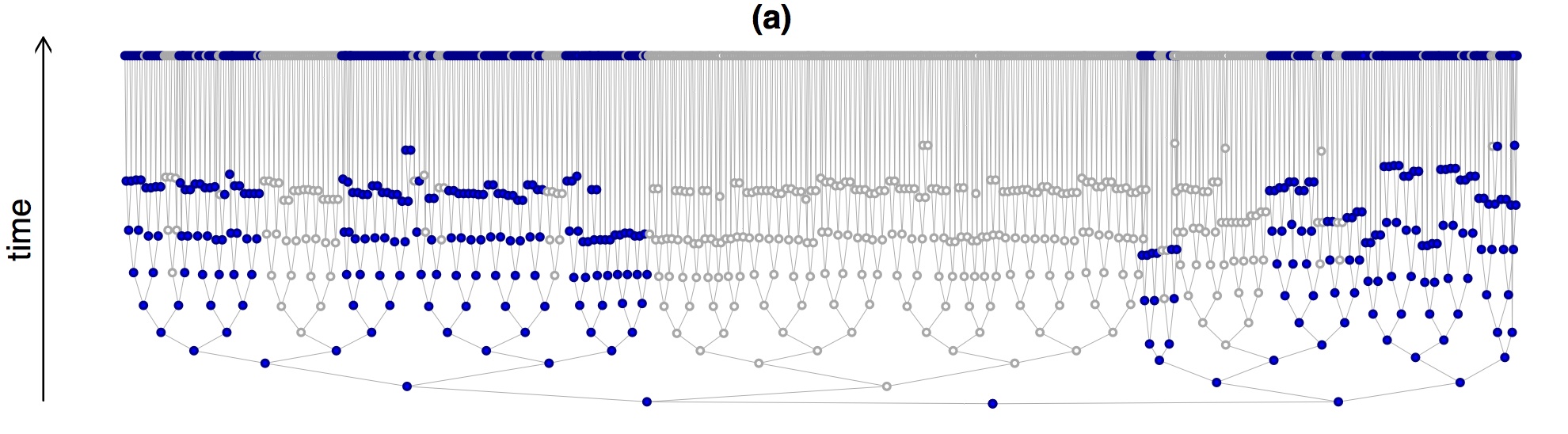
December 2015 - Cellular Barcoding and Erythropoiesis.
All blood cells, red and white, originate from Hematopoietic Stem Cells (HSCs), which reside in bone marrow. The reason, for example, that a bone marrow transplant can be needed after extreme chemotherapy is that it sometimes the case that cancer treatments, as an unwanted side effect, destroys the blood system, and the transplant reseeds it with HSCs from the donor. Given that significance, how, exactly, the reseeded system reboots the blood system has been a significant topic of research in the medical sciences since the discovery of HSCs in the 1960s.
Before becoming a red or white blood cell, of which there are many types, a series of maturation steps takes place where offspring of HSCs progressively acquire specificities. That is, the cells increasingly commit to produce only restricted classes of cell type. When and where that commitment occurs is an important matter as, for example, it identifies where errors that lead to disease must occur and determines where intervention should take place to change outcomes.
The majority of studies determining those commitment steps have been in vitro (i.e. in test tubes), which is the primary distinction of work, "the branching point in erythro-myeloid differentiation" published in Cell. with my colleagues Leïla Perié, Lianne Kok, Rob J. de Boer and Ton N. Schumacher. Leïla designed and performed, with Lianne's aid, a series of experiments using a novel system called cellular barcoding that was developed within the Ton Schumacher's lab. at the Netherlands Cancer Institute. That methodology enables us to observe this commitment in vivo (i.e. within the body). The present work focuses on where in this chain of events the commitment to erythropoiesis, i.e. red blood cell production, takes place. Our analysis of the data from those experiments showed that the commitment is earlier than previously thought.

December 2015 - Quantifying the Computational Security of Multi-User Systems.
The security of many systems is predicated on the following logic: a user selects a string, for example a password, from a list; an inquisitor who knows the list can query each string in turn until gaining access by chancing upon the user's chosen string; the resulting system is deemed to be computationally secure so long as the list of strings is large.
Implicit in that definition is the assumption that the inquisitor knows nothing about the likely nature of the selected string. If instead one assumes that the inquisitor knows the probabilities with which strings are selected, then the number of queries required to identify a stochastically selected string, and the quantification of computational security, becomes substantially more involved. In the mid 1990s, James Massey and Erdal Arikan initiated a programme of research on the susceptibility of such systems to brute for interrogation, dubbed Guesswork, to which from we have been contributing in recent years with collaborators MIT and, more recently, Duke University.
With my my recently graduated Ph.D. student, Mark Christiansen, my colleague Muriel Médard (MIT) and her recently graduated student, Flávio du Pin Calmon (now IBM Research), our most recent contribution has just been published in IEEE Transactions on Information Theory. In it, we address a natural extension in this investigation of brute force searching: the quantification for multi-user systems. We were motivated by both classical systems, such as the brute force entry to a multi-user computer where the inquisitor need only compromise a single account, as well as modern distributed storage services where coded data is kept at distinct sites in a way where, due to coding redundancy, several, but not all, servers need to be compromised to access the content.
The results establish that from an inquisitor's point of view there is a law of diminishing returns as the number of potentially hacked accounts increases. From a designer's point of view, coming full circle to Massey's original observation that Shannon entropy has little quantitative relationship to how hard it is to guess a single string, Shannon entropy plays a fundamental role in the multi-user setting.
November 2015 - Noise in Forensic Analysis of DNA
When cells are taken from a crime scene, a substantial amount of processing is necessary before human identification can be determined. This processing results in both noise and artifacts being added to the measured signal. In a paper published in Forensic Science International: Genetics, my colleagues Ullrich Mönich (now TU Munich), Muriel Médard (MIT), Viveck Cadambe (now Penn. State U.), Lauren Alfonse (BU) and Catherine Grgicak (BU) and I characterize the most fundamental element of this process, baseline noise, for circumstances in which there is little initial DNA, as often occurs with crime-scene samples. Historically, this noise has been modeled as Gaussian when present, but the data generated by Lauren and Catherine does not support that hypothesis and instead suggests that a heavy-tailed distribution is more appropriate. This has ramifications for how noise should be treated in continuous models or how filters should be calibrated by crime-labs.June 2015 - Network Coding, Constraint Satisfaction, Guesswork, and Network Infusion.
Three pieces of work on quite distinct topics will be presented at international conferences this month, one of which is a prize-winner.Ying Cui will present one in London, which has won the best paper award of the Communication Theory Symposium and an overall best paper award, at the IEEE International Conference on Communications. It is a contribution to on Network Coding, relating it to constraint satisfaction problems and was written with collaborators Muriel Médard (MIT), Edmund Yeh (Northeastern) and Doug Leith (Trinity College Dublin),
Ahmad Beirami (Duke & MIT) will present another, written with Robert Calderbank (Duke), myself and Muriel Médard (MIT) at the IEEE Symposium on Information Theory in Hong Kong. The work is an investigation of computational security subject to source constraints.
The last, but certainly not least, will be presented Soheil Feizi (MIT) at the International Conference on Computational Social Science in Finland. It is based on work in progress with myself, Manolis Kellis (MIT) and Muriel Médard (MIT). It forms our contribution the question of how you identify the source, or sources, of an epidemic, an idea or a social network application. Soheil presented an earlier version of the work at RECOMB/ISCB Conference on Regulatory and Systems Genomics in San Diego in November 2014.May 2015 - Information Theory and Cryptography.
In 1949, Claude Shannon published one of his opuses, Communication Theory of Secrecy Systems, in which he introduced a framework for a mathematical theory of cryptography. At a workshop of the 8th International Conference on Information-Theoretic Security (ICITS), Flávio du Pin Calmon presented a piece entitled Revisiting the Shannon Theory approach to cryptography written with Mayank Varia, Muriel Médard, Mark M. Christiansen, myself, Linda Zeger and Joao Barros, revisiting the topic.November 2014 - An Algebra for Immune Responses
Lymphocytes, the key players in an adaptive immune response, have long since been known to need to receive multiple signals to mount an effective defense. That discovery led to the two signal theory of T cell activation. The principle being that two independent signals, antigen followed by costimulation, ensure that lymphocyte expansion is only initiated in response to genuine infection. Redundancy in this second signal has long since left a conundrum in the field.A paper published today in the journal Science brings this theory up to date with a quantitative edge. The work was led by the Walter and Eliza Hall Institute of Medical Research's Philip D. Hodgkin and Susanne Heinzel, driven by Phil's Ph.D. student Julia M. Marchingo, in collaboration with Hodgkin lab. members Andrey Kan, Robyn M. Sutherland, Cameron J. Wellard, Mark R. Dowling, two other WEHI lab. heads Gabrielle T. Belz and Andrew M. Lew, and myself. While the signaling of antigen, costimulation and cytokines are complex and involved, clever experimentation in concert with computer modelling and mathematics revealed a simple additive algebra of T cell expansion; one that puts the old conundrum to bed and will, hopefully, provide a quantitative paradigm for therapeutically manipulating immune response strength.

September 2014 - Randomness, Determinism and Immune Responses
One of the difficult challenges in science is doing justice while framing other people's hypotheses. Steven L. Reiner, one of the main instigators behind investigating the role of asymmetric cell division in immunology, and William C. Adams have neatly laid out a deterministic description of adaptive immune responses in a fascinating opinion piece published in Nature Reviews Immunology. My Australian colleagues Philip D. Hodgkin, one of the main proponents of stochastic processes in immunology, Mark R. Dowling and I have written a brief response in defense of randomness in correspondence to the same journal. The community has not yet acquired the data required to answer how deterministic and stochastic processes interleave to build the complete immune response, but with so many different groups designing experiments and doing analysis based on distinct hypotheses, we're all looking forward to the time when that resolution occurs.July 2014 - Forensic Analysis of DNA
The basis of a DNA fingerprint is the measurement of the number of repeats of microsatellites, short repeated sequenes of of DNA, at various locations in the genome. While the combination of given numbers of repeats are (largely) unique for individuals, in forensics applications often the number of repeats cannot be measured precisely. This occurs as the amount of DNA at a crime scene can be small, requiring amplification before the measurement takes place, which can create false signals or missing data, or the sample itself may be made up of a composite from several individuals in which case one gets a combined, mixed measurement.My colleagues Catherine Grgicak, a Forensic Scientist at Boston University, Desmond Lun, a Computer Scientist at Rutgers, and Muriel Médard, an Electronic Engineer at MIT, have been working in an interdisciplinary team, which I have recently joined, to carefully investigate this topic. The first piece of work I have been involved in is a paper to be presented at Asilomar in November 2014 entitled A signal model for forensic DNA mixtures by members of Catherine and Muriel's labs: Ullrich J. Mönich, Catherine Grgicak, Viveck Cadambe, Jason Yonglin Wu, Genevieve Wellner, Ken Duffy and Muriel Médard. The paper details fundamental statistics of DNA fingerprints created by standard forensics techniques in controlled circumstances in an attempt to build a descriptive model of noise generated in the process.
February 2014 - Cellular Barcoding and Inferring Lineage Pathways
There are many instances where one would wish to know the familial relationships of cells, to be able identify those that came from the same parent. For example, the purpose of a bone-marrow transplant is to place new hematopoietic stem cells into to the recipient so that their blood system can be rebuilt. Does each stem cell contribute equally to this rebuilding? Does chance play a role? Are some stem cells specialized? These are all questions that can only be tackled if one can identify cells from the same progenitor.Cellular barcoding, which Ton N. Schumacher's lab. has been at the forefront of developing, is an experimental method in which small pieces of non-functional DNA into otherwise identical cells. Each small piece, a barcode, can then be found in the progeny of that cell and so those in a common family can be identified. This technique has led to many high-profile results in the past year in everything from immunology, cancer and blood system development, including several from Ton's lab.
In a data analysis contribution to the Cellular Barcoding technique, my colleagues Leïla Perié, Philip D. Hodgkin, Shalin H. Naik, Ton N. Schumacher, Rob J. de Boer, and I have published a paper in Cell Reports that develops a mathematical framework for the analysis of data that comes from cellular barcoding experiments. The framework is designed to draw inferences about the compatibility of potential lineage pathways with the data. As an exemplar, we study data from the blood system taken from one of these high-profile results, published in Nature by S. H. Naik, L. Perié, E. Swart, C. Gerlach, N. van Rooij, R. J. de Boer and Ton N. Schumacher. The analysis suggests the classical model of hematopoiesis is not consistent with the data and, inspired by in vitro deductions from the 80s, we propose an alternate lineage pathway that is consistent.
January 2014 - Integrated Random Walks and Large Busy Periods
Ever since an elegant paper by Venkat Anantharam in 1989, it's been known that the way a random walk becomes large, or how a big queue builds up, is by a piecewise linear path. The question of how a large integrated random walk, or how a large volume of work done during a busy period of a queue, occurs was first tackled by A. A. Borovkov , O. J. Boxma and Z. Palmowski in a paper in 2003.In a paper to appear in Stochastic Systems, Sean Meyn and I establish that the most likely path to a large busy period is concave in general and, remarkably, has a simple form, following a rescaled, upside-down version of the scaled cumulant generating function. Moreover, the path starts and ends in the same fashion as the most likely path to a long queue as identified by Ayalvadi J. Ganesh and Neil O'Connell in 2002.
For 10 billion simulated paths with i.i.d. Gaussian increments, below is a graph of the random walk that hits the highest height and the prediction from Ganesh and O'Connell conditioned on the height. Also plotted is the largest integrated random walk and our prediction conditioned on the area under the curve. The predicted paths start and end with the same slope, but diverge in between.
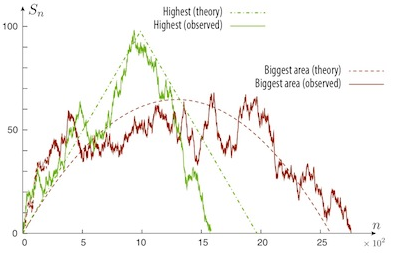
November 2013 - Guesswork, Wiretap and Erasure Channels
Devices such as door-card readers transmit passwords over wireless channels. Unintended receivers, eavesdroppers, are typically distant and observe the communication over a noisy channel that distorts it. How hard is it for the eavesdropper to fill in the erased gaps and how does it depend on the properties of the noisy channel?Complementing earlier work with a distinct interpretation of guesswork and wiretap channels by Neri Merhav and Erdal Arikan, as well as Manjesh Kumar Hanawal and Rajesh Sundaresan, this is a subject that with our collaborators, Flávio du Pin Calmon and Muriel Médard at MIT, Mark Christiansen and I had a paper on at this year's Asilomar Conference on Signals, Systems & Computers. The main observation is that the average noise on the channel is not the determining factor in how difficult the task is for the eavesdropper, but instead another average of the noise, a moment, that is determined again by Rényi entropy.
October 2013 - Limits of Statistical Inference
A paper, which was driven by our collaborators, Flávio du Pin Calmon at MIT and Mayank Varia, at the Lincoln Lab., was presented by Muriel Médard at this year's Allerton conference Communication, Control, and Computing. The work establishes bounds on how much can be inferred about a function of a random variable's value, if only a noisy observation of the variable is available.September 2013 - Constraint Satisfaction and Rate Control
IEEE/ACM Transactions on Networking papers are like buses in that you wait for a long time for one and then two come at once. The work on Decentralized Constraint Satisfaction that is mentioned below has appeared in the August issue of the journal. The final paper from my ex-student Kaidi Huang's doctoral thesis also appears in that issue. The work, performed in collaboration with my Hamilton Institute colleague David Malone, addresses a practical problem in wireless local area networks: rate control. When transmitting packets, WiFi cards must select a physical-layer rate at which to transmit. In principle, the faster the rate, the less robust the transmission is to noise. For the poorly engineered rates of standard WiFi, this is not strictly true as Kaidi, David and I demonstrated in a paper published in IEEE Transactions on Wireless Communications in 2011. In the present work, we proposed a rate adaption scheme based on opportunity cost and Bayesian decision making that was, demonstrably thanks to hard work of Kaidi and David, implementable on standard hardware and outperforms the standard algorithms.July 2013 - Information Theory, Guesswork & Computational Security
The late Kai Lai Chung purportedly said that there's only one theorem in Information Theory: the Asymptotic Equipartition Property. At the 2013 IEEE International Symposium on Information Theory, Mark Christiansen presented a preliminary version of a surprising result that we established in collaboration with our friends from M.I.T., Flávio du Pin Calmon and Muriel Médard. Namely, despite the fact that everything within a Typical Set has, by construction, approximately equal probability, it is exponentially easier as a function of word length to guess a word chosen from a Typical Set than a naive application of the AEP would have you deduce. This has ramifications for physical layer security.January 2013 - Guesswork & Information Theory
How hard is it to guess someone's password? More importantly, how do you measure how hard it is to guess someone's password? It's a question that has recieved less attention than one would have expected. The recently deceased information theorist James Massey published a fascinating one page paper at ISIT on this question in 1994 and it is the topic of a paper that my student, Mark Christiansen, and I have had published in IEEE Transactions on Information Theory. Building on beautiful work of others, which began with Erdal Arikan and included a contribution from my Hamilton Institute colleague David Malone, that identified Rényi entropy as a key player in the quantification of guesswork, in the article we provide a direct estimate on the likelihood that it takes a given number of guesses to guess a randomly selected object.January 2013 - Networking & Medium Access Control
There are two fundamental paradigms for sharing a resource such as wireless medium. One can empower a centralized controller who gathers everyone's requirements and sets out a schedule of who gets access to the resource when. This system is used, for example, for speakers at every conference and in cell phone networks. The alternate system is to not be prescriptive, but to listen before speaking and when the medium becomes silent, randomly interject. This is used, for example, in human conversation as well forming the basis for the standard access method, IEEE 802.11, for WiFi networks.Both of these systems have advantages and disadvantages. If one knows exactly what who needs what resources and can agree on a central party to adjudicate, the former is most efficient. If one is uncertain about the number of users and their requirements, the latter is more robust to this uncertainty, but suffers from collisions - people talking over one another and thus wasting resources.
In a paper that has been published in the journal Wireless Networks, written with my ex-student Minyu Fang, and colleagues at the Hamilton Institute David Malone and Douglas J. Leith, we investigate a system that has the best of both worlds: a decentralized stochastic algorithm is used to obtain a collision-free schedule.
October 2012 - Constraint Satisfaction Problems
Is it possible to find a global solution to a problem, with everyone only having local information and being unable to seethe global picture? That's the topic of a paper that my colleagues, the French mathematician Charles Bordenave
and Douglas J. Leith, my Scottish engineering colleague from the Hamilton Institute, address in a paper that has
been accepted for publication in IEEE/ACM Transactions on Networking. Two undergraduates, TCD's John Roche
and National University of Ireland Maynooth's Tony Poole, created java applets that illustrate how the proposed approach, which provably works, solves problems. They also illustrate that, in this setting, it is easier to agree to disagree than it is to find a consensus.
September 2012 - Immunology & Cell Biology
My Australian immunology collaborator Philip Hodgkin and I had a paper published in Trends in Cell Biology. It expands on the hypothesis we employed in our earlier 2012 paper, published in Science, to explain cell fate diversity. Artwork based on the paper, illustrated by WEHI's Jie H. S. Zhou, was used for the edition's cover:

January 2012 - Immunology & Cellular Biology
With colleagues from the Walter and Eliza Hall Institute of Medical Research led by Philip D. Hodgkin, I had a paper published in Science. The piece analyses data from an experiment that took my Australian colleagues, particularly Mark Dowling and John Markham, four years to perfect. It enabled us to directly observe the times at which an important class of cells in the immune system, the antibody secreting B lymphocytes, make decisions on when and how to combat a pathogen. Despite the data looking immensely complex, it is consistent with a remarkably simple, holistic hypothesis.

Publications
Drafts-
Estimating large deviation rate functions.
Ken R. Duffy, and Brendan D. Williamson. -
Hiding symbols and functions: new metrics and constructions for Information-Theoretic Security.
Flávio du Pin Calmon, Muriel Médard, Mayank Varia, Ken R. Duffy, Mark M. Christiansen and Linda Zeger.
Journal papers
-
2018
-
Network Infusion to infer information sources in networks.
Soheil Feizi, Muriel Médard, Gerald Quon, Manolis Kellis and Ken R. Duffy.
IEEE Transactions on Network Science and Engineering, to appear.
-
Proliferation dynamics of acute myeloid leukaemia and haematopoietic
progenitors competing for bone marrow space.
Olufolake Akinduro, Tom S. Weber, Heather Ang, Myriam Haltalli, Nicola Ruvio, Delfim Duarte, N. M. Rashidi, Edwin D. Hawkins, Ken R. Duffy and Cristina Lo Celso.
Nature Communications, 9, 519, 2018. -
Inhibition of endosteal vascular niche remodeling rescues hematopoietic
stem cell loss in AML.
Delfim Duarte, Edwin D. Hawkins, Olufolake Akinduro, Heather Ang, Katia De Filippo, Isabella Y. Kong, Myriam L. R. Haltalli, Nicola Ruivo, Lenny Straszkowski, Stephin J. Vervoort, Catriona McLean, Tom S. Weber, Reema Khorshed, Chiara Pirillo, Andrew Wei, Saravana K. Ramasamy, Anjali P. Kusumbe, Ken R. Duffy, Ralph H. Adams, Louise E. Purton, Leo M. Carlin and Cristina Lo Celso.
Cell Stem Cell, 22 (1), 64--77, 2018. -
A large-scale dataset of single and mixed-source short tandem repeat profiles to inform human identification strategies: PROVEDIt
Lauren E. Alfonse, Amanda D. Garrett, Desmond Lun, Ken R. Duffy and Catherine Grgicak.
Forensic Science International: Genetics, 32, 62-70, 2018. -
Production of high-fidelity electropherograms results in improved
and consistent DNA interpretation: standardizing the forensic
validation process
Kelsey Peters, Harish Swaminathan, Ken R. Duffy, Desmond Lun, Jennifer Sheehan, and Catherine Grgicak.
Forensic Science International: Genetics, 31, 160-170, 2017. -
A linear network code construction for general integer connections
based on the constraint satisfaction problem.
Ying Cui, Muriel Médard, Edmund Yeh, Douglas J. Leith, Fan Li, and Ken R. Duffy.
IEEE/ACM Transactions on Networking, 25(6), 3441-3454, 2017. -
Network maximal correlation.
Soheil Feizi, Ali Makhdoumi, Ken R. Duffy, Manolis Kellis and Muriel Médard.
IEEE Transactions on Network Science and Engineering, 4(4), 229-247, 2017. -
Principal inertia components and applications.
Flávio du Pin Calmon, Ali Makhdoumi, Muriel Médard, Mayank Varia, Mark M. Christiansen and Ken R. Duffy,
IEEE Transactions on Information Theory, 63(8), 5011-5038, 2017. -
Exploring forensic STR signal in the single- and multi-copy number regimes: deductions from an in silico model of the entire DNA laboratory process.
Ken R. Duffy, Neil Gurram, Kelsey Peters, Genevieve Wellner, and Catherine Grgicak.
Electrophoresis, 38(6), 855–868, 2017. -
T cell stimuli independently sum to regulate an inherited clonal division fate.
Julia M. Marchingo, Giulio Prevedello, Andrey Kan, Susanne Heinzel, Philip D. Hodgkin (†) and Ken R. Duffy (†).
Nature Communications, 7, 13540, 2016.
-
Re-tracing the in vivo hematopoietic tree using single cell methods.
Leïla Perié and Ken R. Duffy.
FEBS Letters, 590 (22), 4068-4083, 2016. -
Site-specific recombinatorics: in situ cellular barcoding with the Cre Lox system.
Tom S. Weber, Mark Dukes, Denise C. Miles, Stefan P. Glaser, Shalin H. Naik, and Ken R. Duffy.
BMC Systems Biology, 10:42, 2016. -
Inferring average generation via division-linked labeling.
Tom S. Weber, Leïla Perié and Ken R. Duffy.
Journal of Mathematical Biology, 73 (2), 491-523, 2016. -
The branching point in erythro-myeloid differentiation.
Leïla Perié, Ken R. Duffy, Lianne Kok, Rob J. de Boer and Ton N. Schumacher.
Cell, 163 (7), 1655-1662, 2015.
-
Multi-user guesswork and brute force security.
Mark M. Christiansen, Ken R. Duffy, Flávio du Pin Calmon and Muriel Médard.
IEEE Transactions on Information Theory, 61 (12), 6876-6886, 2015.
-
Probabilistic characterisation of baseline noise in STR profiles.
Ullrich J. Mönich, Ken R. Duffy, Muriel Médard, Viveck Cadambe, Lauren E. Alfonse and Catherine Grgicak.
Forensic Science International: Genetics, 19, 107-122, 2015. -
Antigen affinity, costimulation, and cytokine inputs sum linearly
to amplify T cell expansion.
Julia M. Marchingo, Andrey Kan, Robyn M. Sutherland, Ken R. Duffy, Cameron J. Wellard, Gabrielle T. Belz, Andrew M. Lew, Mark R. Dowling, Susanne Heinzel, Philip D. Hodgkin.
Science, 346 (6213), 1123-1127, 2014.
-
Determining lineage pathways from cellular barcoding experiments.
Leïla Periè, Philip D. Hodgkin, Shalin H. Naik, Ton N. Schumacher, Rob J. de Boer and Ken R. Duffy.
Cell Reports 6 (4), 617-624, 2014.
-
Large deviation asymptotics for busy periods.
Ken R. Duffy and Sean P. Meyn.
Stochastic Systems 4 (1), 300-319, 2014.
2013
-
Decentralized constraint satisfaction.
Ken R. Duffy, Charles Bordenave and Douglas J. Leith.
IEEE/ACM Transactions on Networking, 21 (4), 1298-1308, 2013.
-
H-RCA: 802.11 collision-aware rate control.
K. D. Huang, Ken R. Duffy and David Malone.
IEEE/ACM Transactions on Networking, 21 (4), 1021-1034, 2013.
-
Guesswork, large deviations and Shannon entropy.
Mark M. Christiansen and Ken R. Duffy.
IEEE Transactions on Information Theory, 59 (2), 796-802, 2013.
-
Decentralised learning MACs for collision-free
access in WLANs.
Minyu Fang, David Malone, Ken R. Duffy and Douglas J. Leith.
Wireless Networks, 19 (1), 83-98, 2013.
2012
-
Intracellular competition for fates in the immune system.
Ken R. Duffy and Philip D. Hodgkin.
Trends in Cell Biology, 22 (9), 457-464, 2012.
- Activation-induced B cell fates are selected by intracellular stochastic competition.
Ken R. Duffy, Cameron J. Wellard, John F. Markham, Jie H. S. Zhou, Ross Holmberg,
Edwin D. Hawkins, Jhagvaral Hasbold, Mark R. Dowling and Philip D. Hodgkin.
Science, 335 (6066), 338-341, 2012.
2011
-
Sample path large deviations of Poisson shot noise with heavy tail
semi-exponential distributions.
Ken R. Duffy and Giovanni Luca Torrisi.
Journal of Applied Probability, 48 (3), 688-698, 2011. -
Estimating Loynes' exponent.
Ken R. Duffy and Sean P. Meyn.
Queueing Systems: Theory & Applications, 68 (3-4), 285-293, 2011. -
The 802.11g 11 Mb/s rate is more robust than 6 Mb/s.
K. D. Huang, David Malone and Ken R. Duffy.
IEEE Transactions on Wireless Communcations, 10 (4), 1015-1020, 2011. -
Sample path large deviations for order statistics.
Ken R. Duffy, Claudio Macci and Giovanni Luca Torrisi.
Journal of Applied Probability, 48 (1), 238-257, 2011. -
On the large deviations of a class of modulated additive
processes.
Ken R. Duffy, Claudio Macci and Giovanni Luca Torrisi.
ESAIM: Probability and Statistics, 2011 (15), 83-109, 2011.
2010
-
Most likely paths to error when estimating the mean of a reflected
random walk.
Ken R. Duffy and Sean P. Meyn.
Performance Evaluation, 67 (12), 1290-1303, 2010. -
On the validity of IEEE 802.11 MAC modeling hypotheses.
K. D. Huang, Ken R. Duffy and David Malone.
IEEE/ACM Transactions on Networking, 18 (6), 1935-1948, 2010. -
Mean field Markov models of wireless local area networks.
Ken R. Duffy.
Markov Processes and Related Fields, 16 (2), 295-328, 2010. -
A minimum of two distinct heritable factors are required to explain
correlation structures in proliferating lymphocytes.
John F. Markham, Cameron J. Wellard, Edwin D. Hawkins, Ken R. Duffy and Philip D. Hodgkin.
Journal of the Royal Society Interface, 7 (48), 1049-1059, 2010.
-
Log-convexity of rate region in 802.11e WLANs.
Douglas J. Leith, Vijay G. Subramanian and Ken R. Duffy.
IEEE Communications Letters, 14 (1), 57-59, 2010.
2009
-
Existence and uniqueness of fair rate allocations
in lossy wireless networks.
Vijay G. Subramanian, Ken R. Duffy and Douglas J. Leith.
IEEE Transactions on Wireless Communications, 8 (7), 3401-3406, 2009. -
On a buffering hypothesis in 802.11 analytic models.
K. D. Huang and Ken R. Duffy
IEEE Communications Letters, 13 (5), 312-314, 2009. -
On the impact of correlation between collaterally consanguineous cells
on lymphocyte population dynamics.
Ken R. Duffy and Vijay G. Subramanian.
Journal of Mathematical Biology, 59 (2), 255-285, 2009.
2008
-
Complexity analysis of a decentralised graph colouring algorithm.
Ken R. Duffy, Neil O'Connell and Artem Sapozhnikov.
Information Processing Letters, 107 (2), 60-63, 2008. -
The large deviation principle for the on/off Weibull sojourn process.
Ken R. Duffy and Artem Sapozhnikov.
Journal of Applied Probability, 45 (1), 107-117, 2008. -
Determining the expected variability of immune responses using the Cyton Model.
Vijay G. Subramanian, Ken R. Duffy, Marian L. Turner and Philip D. Hodgkin.
Journal of Mathematical Biology, 56 (6), 861-892, 2008. -
Logarithmic asymptotics for a single-server processing distinguishable sources.
Ken R. Duffy and David Malone.
Mathematical Methods of Operations Research, 68 (3), 509-537, 2008.
2007
-
Loss aversion, large deviation preferences and optimal portfolio weights
for some classes of return processes.
Ken Duffy, Olena Lobunets and Yuri Suhov.
Physica A, 378 (2), 408-422, 2007. -
Modeling the impact of buffering on 802.11.
Ken Duffy and Ayalvadi J. Ganesh.
IEEE Communications Letters, 11 (2), 219-221, 2007. -
Ambiguities in estimates of critical exponents for long-range dependent processes.
Ken Duffy, Christopher King and David Malone.
Physica A, 377 (1), 43-52, 2007. -
Some remarks on LD plots for heavy-tailed traffic.
David Malone, Ken Duffy and Christopher King.
ACM SIGCOMM Computer Communications Review, 37 (1), 41-42, 2007. -
Using estimated entropy in a queueing system with dynamic routing.
Ken Duffy, Eugene A. Pechersky, Yuri M. Suhov and Nikita D. Vvedenskaya.
Markov Processes and Related Fields, 13 (1-2), 57-84, 2007. -
Modeling the 802.11 distributed coordination function in
non-saturated heterogeneous conditions.
David Malone, Ken Duffy and Douglas J. Leith.
IEEE/ACM Transactions on Networking, 15 (1), 159-172, 2007.
2006
-
Modeling 802.11 Mesh Networks.
Ken Duffy, Douglas J. Leith, Tianji Li and David Malone.
IEEE Communications Letters 10 (8), 635-637, 2006.
2005
-
How to estimate a cumulative process's rate-function.
Ken Duffy and Anthony P. Metcalfe.
Journal of Applied Probability 42 (4), 1044-1052, 2005. -
Modeling the 802.11 distributed coordination function in
non-saturated conditions.
Ken Duffy, David Malone and Douglas J. Leith.
IEEE Communications Letters, 9 (8), 715-717, 2005. -
The large deviations of estimating rate-functions.
Ken Duffy and Anthony P. Metcalfe.
Journal of Applied Probability 42 (1), 267-274, 2005.
2004
-
Some useful functions for functional large deviations.
Ken Duffy and Mark Rodgers-Lee.
Stochastics and Stochastics Reports 76 (3), 267-279, 2004. -
Logarithmic asymptotics for unserved messages at a FIFO.
Ken Duffy and Wayne G. Sullivan.
Markov Processes and Related Fields 10 (1), 175-189, 2004. -
On Knuth's generalization of Banach's matchbox problem.
Ken Duffy and W. M. B. Dukes.
Mathematical Proceedings of the Royal Irish Academy 104A, 107-118, 2004.
2003
-
Logarithmic asymptotics for the supremum of a stochastic process.
Ken Duffy, John T. Lewis and Wayne G. Sullivan.
Annals of Applied Probability 13:2, 430-445, 2003.
2000
-
Distribution-free confidence intervals for measurements of
effective bandwidth.
Laszlo Gyorfi, Andras Racz, Ken Duffy, John T. Lewis and Fergal Toomey.
Journal of Applied Probability 37:1, 224-235, 2000.
2017
2016
2015
2014
Conference papers
-
Guessing noise, not code-words.
Ken R. Duffy, Jiange Li, and Muriel Médard.
IEEE International Symposium on Information Theory, 17th-22nd June 2018, Colorado, USA.
-
Guesswork subject to a total entropy budget.
Arman Rezaee, Ahmad Beirami, Ali Makhdoumi, Muriel Médard and Ken R. Duffy.
55th Allerton Conference on Communication, Control, and Computing, 4th-6th October 2017, Illinois, USA.
-
A geometric perspective on guesswork.
Ahmad Beirami, Robert Calderbank, Mark M. Christiansen, Ken R. Duffy Ali Makhdoumi, and Muriel Médard.
53rd Allerton Conference on Communication, Control, and Computing, 30th September - 2nd October 2015, Illinois, USA. -
A linear Network Code construction for general integer connections
based on the Constraint Satisfaction Problem.
Ying Cui, Dhaivat Pandya, Muriel Médard, Edmund Yeh, Douglas J. Leith, and Ken R. Duffy.
IEEE Global Communications Conference, 6th-10th December 2015, San Diego, USA. -
Computational security subject to source constraints, guesswork
and inscrutability.
Ahmad Beirami, Robert Calderbank, Ken R. Duffy and Muriel Médard.
IEEE International Symposium on Information Theory , 14th-19th June 2015, Hong Kong. -
Optimization-based linear Network Coding for general connections
of continuous flows.
Ying Cui, Muriel Médard, Edmund Yeh, Douglas J. Leith, and Ken R. Duffy.
Winner of the best paper award of the Communication Theory Symposium and an overall best paper award.
IEEE International Conference on Communications, 8th-12th June 2015, London, UK.
-
A signal model for forensic DNA mixtures.
Ullrich J. Mönich, Catherine Grgicak, Viveck Cadambe, Jason Yonglin Wu, Genevieve Wellner, Ken Duffy and Muriel Médard.
Asilomar Conference on Signals, Systems & Computers, November 2nd-5th 2014, California, USA. -
Guessing a password over a wireless channel (on the effect of noise
non-uniformity).
Mark M. Christiansen, Ken R. Duffy, Flávio du Pin Calmon and Muriel Médard.
Asilomar Conference on Signals, Systems & Computers, 3rd-6th November, 2013, California, USA. -
Bounds on inference.
Flávio du Pin Calmon, Mayank Varia, Muriel Médard, Mark M. Christiansen, Ken R. Duffy and Stefano Tessaro.
51st Allerton Conference on Communication, Control, and Computing, 2nd-4th October 2013, Illinois, USA. -
Brute force searching, the typical set and Guesswork.
Mark M. Christiansen, Ken R. Duffy, Flávio du Pin Calmon and Muriel Médard.
IEEE International Symposium on Information Theory, 7th-12th July, 2013, Istanbul, Turkey. -
Lists that are smaller than their parts: A coding approach
to tunable secrecy.
Flávio Calmon, Muriel Médard, Linda Zeger, Joao Barros, Mark M. Christiansen and Ken R. Duffy.
50th Annual Allerton Conference on Communication, Control, and Computing, 1st-5th October 2012, Illinois, USA. -
Modeling conservative updates in multi-hash approximate count sketches.
Giuseppe Bianchi, Ken R. Duffy, Douglas J. Leith and Seva Shneer.
24th International Teletraffic Congress, 4-7th September 2012, Krakow, Poland. -
Investigating the validity of IEEE 802.11 MAC modeling hypotheses.
K. D. Huang, Ken R. Duffy, David Malone and Douglas J. Leith.
IEEE PIMRC 15-18th September 2008, Cannes, France. -
Improving fairness in multi-hop mesh networks using 802.11e.
Ken Duffy, Douglas J. Leith, Tianji Li and David Malone.
Resource Allocation in Wireless Networks, 3rd April 2006, Boston, USA. -
Modeling 802.11e for data traffic parameter design.
Peter Clifford, Ken Duffy, John Foy, Douglas J. Leith and David Malone.
WiOPT 2006, 28-37, 3rd-7th April 2006, Boston, USA. -
Modelling 802.11 wireless links.
Ken Duffy, David Malone and Douglas J. Leith.
IEEE Conference on Decision and Control, 6952 - 6957, 12th-15th December 2005, Seville, Spain. -
On improving voice capacity in 802.11 infrastructure networks.
Peter Clifford, Ken Duffy, Douglas J. Leith and David Malone.
IEEE WirelessCom, 214 - 219, 13th-16th June 2005, Maui, Hawaii, USA. -
Modeling the 802.11 distributed coordination function with heterogeneous
finite load.
David Malone, Ken Duffy and Douglas J. Leith.
Resource Allocation in Wireless Networks, 3rd April 2005, Trento, Italy.
-
Why the immune system takes its chances with randomness.
Philip D. Hodgkin, Mark R. Dowling and Ken R. Duffy.
Nature Reviews Immunology, 14 (10), 711, 2014.
Correspondence in response to an interesting opinion piece by Steven L. Reiner and William C. Adams. -
Modelling and simulation analysis of the 802.11/802.11e MAC layers.
Qiang Ni, David Malone, Peter Clifford, Ken Duffy, Douglas Leith and Tianji Li.
Advances in Wireless Networks: Performance Modelling, Analysis and Enhancement,
Nova Science Publishers, edited by Geyong Min, Yi Pan and Pingzhi Fan, 2007. -
John T. Lewis (1932-2004).
Ken Duffy and Joe Pule.
Markov Processes and Related Fields, 13 (1-2), 3-19, 2007.
Research Funding
Recent Funding
-
"Establishing exclusion criteria and the significance of inclusion:
developing tools for the interpretation of complex DNA mixtures",
2015-2017, National Institute of Justice (USA), sub-contractor. PIs: Catherine Grgicak (Boston U.) and Desmond Lun (Rutgers). -
"Quantitative analysis of immune cell fate: stochastic competition
and censorship",
2013-2017, Science Foundation Ireland Investigator Grant. -
"Quantitative T cell immunology",
2013-2017, European Union Marie Curie FP7 ITN, co-PI with 16 other institutions, including industry, with co-ordinating co-PI Grant Lythe (Leeds). -
"Indo-European research network in mathematics for health and disease",
2013-2017, European Union Marie Curie FP7 IRSES, co-PI with five other institution, co-ordinating co-PI Carmen Molina-Paris (Leeds), -
"Single cell lineage tracing to understand hematopoietic development and differentiation",
2012-2015, Human Frontier Science Program Research Grant, co-PI with Andrew Cohen (Drexel), Philip Hodgkin (WEHI) and Ton Schumacher (NKI). -
"FLAVIA: Flexible architecture for virtualizable future wireless internet access"
2010-2013, European Union Framework 7 Strep Grant, collaborator, ten partner academic and industry grant led by Giuseppe Bianchi (U. Rome), and locally by David Malone (Maynooth University). -
"Mathematical modelling of lymphocyte proliferation and differentiation
during an adaptive immune response",
2009, Science Foundation Ireland Short-term Travel Fellowship to visit Philip Hodgkin (WEHI). -
"Using 802.11 medium access control layer measurements to
understand and improve network performance",
2007-2010, Science Foundation Ireland Research Frontiers Programme, PI, with David Malone (Maynooth University) as co-PI.
Students
The group on sabbatical at MIT in 2016 (L-R: Gianfelice, Harry, Alex, Giulio, Neil, myself and Tom):

Current graduate-students
- Gianfelice Meli.
- Harry Tideswell.
- Giulio Prevedello.
Past graduate-students
-
Alexander Miles, M.Sc. (National University of Ireland Maynooth), 2017.
-
Kelsey Peters M.S. (Boston University Medical School,
2nd supervisor to
Prof. Catherine Grgicak), 2017.
-
Neil Gurram, M.Eng. (Massachusetts Institute of Technology), 2016.
-
Mark Christiansen, Ph.D. (National University of Ireland Maynooth), 2015.
-
Kaidi Huang, Ph.D. (National University of Ireland Maynooth), 2010.
-
Minyu Fang, M.Sc. (National University of Ireland Maynooth), 2010.
- Anthony Paul Metcalfe, M.Sc. (University of Dublin), 2004.
- Mark Rodgers-Lee, M.Sc. (University of Dublin), 2003.
Interns
- Sarah Halpin, Theoretical Physics (University of Dublin), 2017.
- Brendan Williamson, Mathematics (Duke University), 2014.
- Conor Leonard, Mathematics (University College Cork), 2013.
- Brendan Williamson, Mathematics (Dublin City University), 2012.
- John Roche, Mathematics (University of Dublin), 2011.
- Tony Poole, Science (National University of Ireland Maynooth), 2011.
- Joshua Tobin, Mathematics (University of Dublin), 2010.